🔬 Fluorescent In Situ Hybridization (FISH)
Visualizing MAP at the Cellular Level with Precision

What is FISH?
A molecular technique called Fluorescent In Situ Hybridization (FISH) locates and identifies particular genetic sequences within cells or tissues using fluorescently labeled DNA or RNA probes. When it comes to MAP, FISH enables researchers and medical professionals to visually verify that MAP genetic material is present inside biological samples like tissue slices, milk cells, or fecal smears.
FISH is perfect for research and diagnostics because it shows the target's location in the sample, unlike PCR, which amplifies DNA.
The Operation of FISH
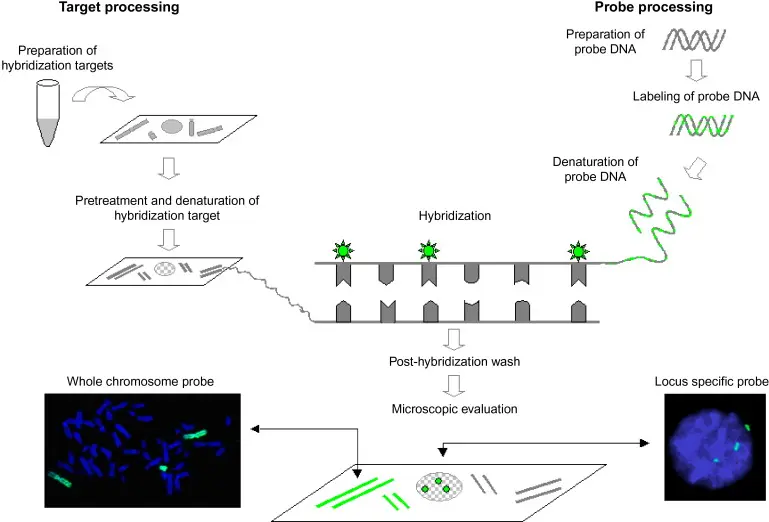
Sample Preparation: To enable probe entry, tissue or cell samples are adhered to a slide and permeabilized.
Probe Hybridization: The sample is mixed with fluorescent DNA or RNA probes that are made to match particular MAP sequences (usually IS900).
Washing and Detection: Probes that are unbound are removed by washing. Positive signals show up as bright dots under a fluorescence microscope, indicating that MAP DNA or RNA is present.
Analysis: It is possible to thoroughly examine the spatial distribution of MAP, both inside host cells and even within particular tissue layers.
Why Use FISH for MAP?
FISH offers unique advantages in MAP detection:
- Spatial Localization: MAP can be visualized inside macrophages, epithelial cells, or tissue lesions.
- Direct Evidence: Unlike culture methods that take months, FISH provides same-day results from clinical samples.
- Gene-Specific Targeting: Probes can target not only IS900, but also other genes like F57 or 16S rRNA, allowing for specificity and flexibility.
- Dual Staining: FISH can be combined with host markers to study host-pathogen interactions in situ.
MAP detection in pure culture, sheep, and human tissue using FISH hybridization of P90CorrProbe :
Uses in MAP Diagnostics and Research
Histopathological Confirmation: FISH aids in confirming the existence of MAP in ruminant tissues that have been formalin-fixed paraffin-embedded (FFPE).
Milk Quality Control: To ensure food safety, find MAP DNA in milk somatic cells.
Investigate the pathophysiology of MAP by examining its interactions with immune cells or the gut lining in animals and experimental models.
Co-detection: To help with differential diagnosis, FISH can be multiplexed with probes for additional bacterial pathogens or mycobacteria.
Advantages
- Specific and highly visual
- Works directly on preserved samples
- Enables co-localization with host markers
- Detects both DNA and RNA (for active infection profiling)
The Future of FISH in MAP Detection
FISH is developing into a high-resolution diagnostic and research platform with automation, improved image analysis, and multiplexing technologies. It is an essential tool for researching cellular host-pathogen interactions and chronic infections like Johne's disease because of its capacity to offer spatial context.